Workflow: QuantSeq 3' mRNA-Seq single-read
### Pipeline for Lexogen's QuantSeq 3' mRNA-Seq Library Prep Kit FWD for Illumina [Lexogen original documentation](https://www.lexogen.com/quantseq-3mrna-sequencing/) * Cost-saving and streamlined globin mRNA depletion during QuantSeq library preparation * Genome-wide analysis of gene expression * Cost-efficient alternative to microarrays and standard RNA-Seq * Down to 100 pg total RNA input * Applicable for low quality and FFPE samples * Single-read sequencing of up to 9,216 samples/lane * Dual indexing and Unique Molecular Identifiers (UMIs) are available ### QuantSeq 3’ mRNA-Seq Library Prep Kit FWD for Illumina The QuantSeq FWD Kit is a library preparation protocol designed to generate Illumina compatible libraries of sequences close to the 3’ end of polyadenylated RNA. QuantSeq FWD contains the Illumina Read 1 linker sequence in the second strand synthesis primer, hence NGS reads are generated towards the poly(A) tail, directly reflecting the mRNA sequence (see workflow). This version is the recommended standard for gene expression analysis. Lexogen furthermore provides a high-throughput version with optional dual indexing (i5 and i7 indices) allowing up to 9,216 samples to be multiplexed in one lane. #### Analysis of Low Input and Low Quality Samples The required input amount of total RNA is as low as 100 pg. QuantSeq is suitable to reproducibly generate libraries from low quality RNA, including FFPE samples. See Fig.1 and 2 for a comparison of two different RNA qualities (FFPE and fresh frozen cryo-block) of the same sample. 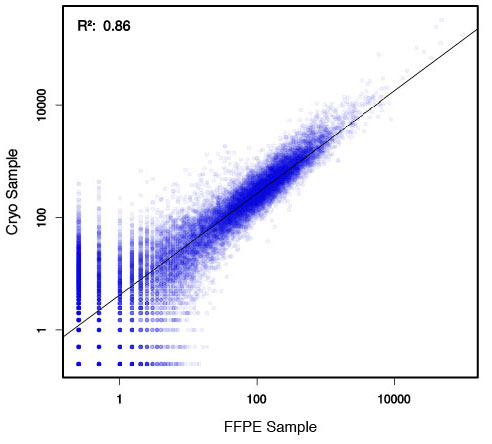 Figure 1 | Correlation of gene counts of FFPE and cryo samples. 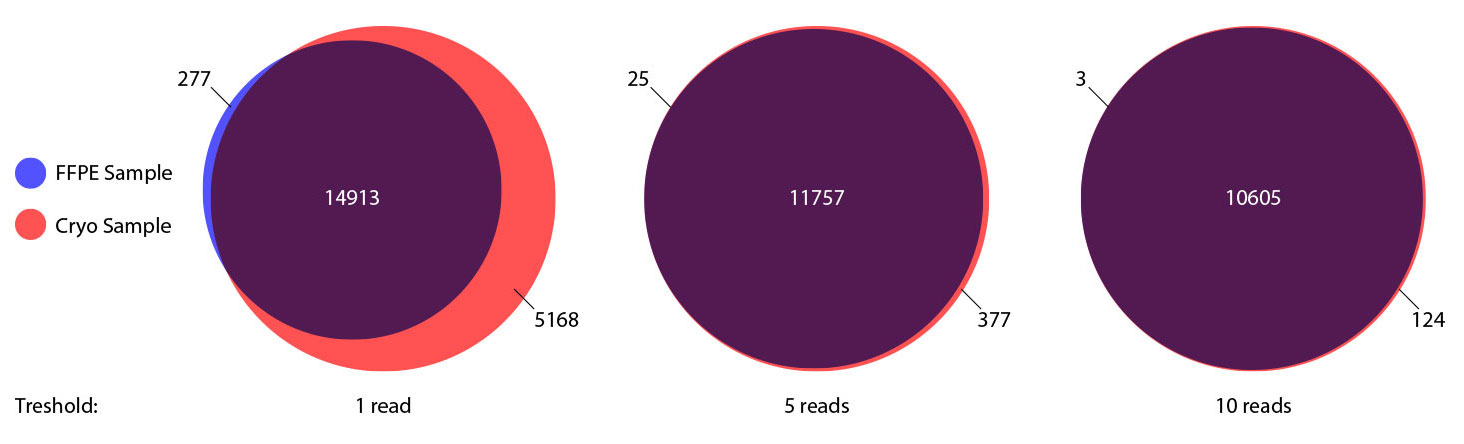 Figure 2 | Venn diagrams of genes detected by QuantSeq at a uniform read depth of 2.5 M reads in FFPE and cryo samples with 1, 5, and 10 reads/gene thresholds. #### Mapping of Transcript End Sites By using longer reads QuantSeq FWD allows to exactly pinpoint the 3’ end of poly(A) RNA (see Fig. 3) and therefore obtain accurate information about the 3’ UTR. 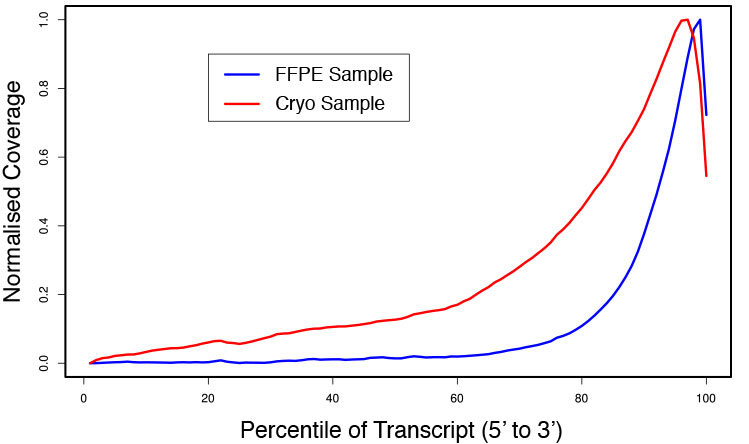 Figure 3 | QuantSeq read coverage versus normalized transcript length of NGS libraries derived from FFPE-RNA (blue) and cryo-preserved RNA (red). ### Current workflow should be used only with the single-end RNA-Seq data. It performs the following steps: 1. Separates UMIes and trims adapters from input FASTQ file 2. Uses ```STAR``` to align reads from input FASTQ file according to the predefined reference indices; generates unsorted BAM file and alignment statistics file 3. Uses ```fastx_quality_stats``` to analyze input FASTQ file and generates quality statistics file 4. Uses ```samtools sort``` and generates coordinate sorted BAM(+BAI) file pair from the unsorted BAM file obtained on the step 2 (after running STAR) 5. Uses ```umi_tools dedup``` and generates final filtered sorted BAM(+BAI) file pair 6. Generates BigWig file on the base of sorted BAM file 7. Maps input FASTQ file to predefined rRNA reference indices using ```bowtie``` to define the level of rRNA contamination; exports resulted statistics to file 8. Calculates isoform expression level for the sorted BAM file and GTF/TAB annotation file using GEEP reads-counting utility; exports results to file
- Selected
- |
- Default Values
- Nested Workflows
- Tools
- Inputs/Outputs
Inputs
| ID | Type | Title | Doc |
|---|---|---|---|
| threads | Integer (Optional) | Number of threads |
Number of threads for those steps that support multi-threading |
| use_umi | Boolean (Optional) | Use UMIs |
Use UMIs (for FWD-UMI libraries) |
| fastq_file | File [FASTQ] | FASTQ input file |
Reads data in a FASTQ format |
| clip_3p_end | Integer (Optional) | Clip from 3p end |
Number of bases to clip from the 3p end |
| clip_5p_end | Integer (Optional) | Clip from 5p end |
Number of bases to clip from the 5p end |
| exclude_chr | String (Optional) | Chromosome to be excluded in rpkm calculation |
Chromosome to be excluded in rpkm calculation |
| annotation_file | File [GTF] | Annotation file |
GTF or TAB-separated annotation file |
| chrom_length_file | File [Textual format] | Chromosome length file |
Chromosome length file |
| annotation_gtf_file | File [GTF] | GTF annotation file |
GTF annotation file |
| star_indices_folder | Directory | STAR indices folder |
Path to STAR generated indices |
| bowtie_indices_folder | Directory | BowTie Ribosomal Indices |
Path to Bowtie generated indices |
Steps
| ID | Runs | Label | Doc |
|---|---|---|---|
| rename |
../tools/rename.cwl
(CommandLineTool)
|
Tool renames `source_file` to `target_filename`. Input `target_filename` should be set as string. If it's a full path, only basename will be used. If BAI file is present, it will be renamed too |
|
| get_stat |
../tools/collect-statistics-rna-quantseq.cwl
(CommandLineTool)
|
Tool processes and combines log files generated by Trimgalore, Bowtie, Samtools and MACS2. |
|
| star_aligner |
../tools/star-alignreads.cwl
(CommandLineTool)
|
Tool runs STAR alignReads. |
|
| bam_to_bigwig |
../tools/bam-bedgraph-bigwig.cwl
(Workflow)
|
Workflow converts input BAM file into bigWig and bedGraph files. |
|
| extract_fastq |
../tools/extract-fastq.cwl
(CommandLineTool)
|
Tool to decompress input FASTQ file(s). If several FASTQ files are provided, they will be concatenated in the order that corresponds to files in input. Bash script's logic: - disable case sensitive glob check - check if root name of input file already include '.fastq' or '.fq' extension. If yes, set DEFAULT_EXT to \"\", otherwise use '.fastq' - check file type, decompress if needed - return 1, if file type is not recognized This script also works of input file doesn't have any extension at all |
|
| bowtie_aligner |
../tools/bowtie-alignreads.cwl
(CommandLineTool)
|
Tool maps input raw reads files to reference genome using Bowtie. |
|
| group_isoforms |
trim-quantseq-mrnaseq-se.cwl#group_isoforms/ca8c12b8-20a3-4fe3-b540-77477e7c557b
(CommandLineTool)
|
||
| umi_tools_dedup |
../tools/umi-tools-dedup.cwl
(CommandLineTool)
|
Deduplicate BAM files based on the first mapping co-ordinate and the UMI attached to the read Only -I, --paired and -S parameters are implemented. |
|
| umisep_cutadapt |
trim-quantseq-mrnaseq-se.cwl#umisep_cutadapt/e7048965-84d6-4b9d-bca9-541725e98664
(CommandLineTool)
|
||
| rpkm_calculation |
../tools/geep.cwl
(CommandLineTool)
|
geep |
Tool calculates RPKM values grouped by isoforms or genes. |
| get_bam_statistics |
../tools/samtools-stats.cwl
(CommandLineTool)
|
Generates statistics for the input BAM file. |
|
| fastx_quality_stats |
../tools/fastx-quality-stats.cwl
(CommandLineTool)
|
Tool calculates statistics on the base of FASTQ file quality scores. If `output_filename` is not provided call function `default_output_filename` to return default output file name generated as `input_file` basename + `.fastxstat` extension. |
|
| samtools_sort_index_1 |
../tools/samtools-sort-index.cwl
(CommandLineTool)
|
Tool to sort and index input BAM/SAM/CRAM.
If input `trigger` is set to `true` or isn't set at all (`true` is used by default), run `samtools sort` and
`samtools index`, return sorted BAM and BAI/CSI index file.
If input `trigger` is set to `false`, return unchanged `sort_input` (BAM/SAM/CRAM) and index (BAI/CSI, if provided in
`secondaryFiles`) files, previously staged into output directory. |
|
| samtools_sort_index_2 |
../tools/samtools-sort-index.cwl
(CommandLineTool)
|
Tool to sort and index input BAM/SAM/CRAM.
If input `trigger` is set to `true` or isn't set at all (`true` is used by default), run `samtools sort` and
`samtools index`, return sorted BAM and BAI/CSI index file.
If input `trigger` is set to `false`, return unchanged `sort_input` (BAM/SAM/CRAM) and index (BAI/CSI, if provided in
`secondaryFiles`) files, previously staged into output directory. |
|
| htseq_count_gene_expression |
../tools/htseq-count.cwl
(CommandLineTool)
|
HTSeq: Analysing high-throughput sequencing data |
For convenience to use in the workflow that sort and index BAM files by coordinate
this tools expects coordinate sorted and indexed BAM file as input. For single-read
dat it won't influence on anything, for paired-end the more memory will be used to
keep reads while looking for their proper pairs (see --max-reads-in-buffer parameter). |
Outputs
| ID | Type | Label | Doc |
|---|---|---|---|
| bigwig | File [bigWig] | BigWig file |
Generated BigWig file |
| bowtie_log | File [Textual format] | Bowtie alignment log |
Bowtie alignment log file |
| rpkm_genes | File [TSV] | raw reads grouped by gene name |
raw reads grouped by gene name |
| bambai_pair | File [BAM] | Coordinate sorted BAM alignment file (+index BAI) |
Coordinate sorted BAM file and BAI index file |
| star_sj_log | File (Optional) [Textual format] | STAR sj log |
STAR SJ.out.tab |
| trim_report | File | cutadapt report |
cutadapt generated log |
| get_stat_log | File (Optional) [YAML] | YAML formatted combined log |
YAML formatted combined log |
| star_out_log | File (Optional) [Textual format] | STAR log out |
STAR Log.out |
| star_final_log | File [Textual format] | STAR final log |
STAR Log.final.out |
| rpkm_common_tss | File [TSV] | raw reads grouped by common TSS |
raw reads grouped by common TSS |
| star_stdout_log | File (Optional) [Textual format] | STAR stdout log |
STAR Log.std.out |
| fastx_statistics | File [Textual format] | FASTQ statistics |
fastx_quality_stats generated FASTQ file quality statistics file |
| get_stat_markdown | File (Optional) [TIDE TXT] | Markdown formatted combined log |
Markdown formatted combined log |
| star_progress_log | File (Optional) [Textual format] | STAR progress log |
STAR Log.progress.out |
| get_formatted_stats | File (Optional) [Textual format] | Bowtie, STAR and GEEP mapping stats |
Processed and combined Bowtie & STAR aligner and GEEP logs |
| bam_statistics_report | File [Textual format] | BAM statistics report |
BAM statistics report (right after alignment and sorting) |
| umi_tools_dedup_stats | File[] (Optional) | umi_tools dedup stats |
umi_tools dedup stats |
| htseq_count_stderr_log | File [Textual format] | HTSeq: stderr log |
HTSeq: stderr log |
| htseq_count_stdout_log | File [Textual format] | HTSeq: stdout log |
HTSeq: stdout log |
| umi_tools_dedup_stderr | File | umi_tools dedup stderr log |
umi_tools dedup stderr log |
| umi_tools_dedup_stdout | File | umi_tools dedup stdout log |
umi_tools dedup stdout log |
| htseq_count_report_file | File [TSV] | HTSeq: read counts grouped by gene_id |
Feature counts from htseq-count grouped by gene_id |
https://w3id.org/cwl/view/git/e0a30aa1ad516dd2ec0e9ce006428964b840daf4/workflows/trim-quantseq-mrnaseq-se.cwl






